ALK_ExonImbalance_SKCM_Analysis
Haider Inam
2/16/2019
Last updated: 2019-05-21
Checks: 5 1
Knit directory: pair_con_select/
This reproducible R Markdown analysis was created with workflowr (version 1.3.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of the R Markdown file created these results, you’ll want to first commit it to the Git repo. If you’re still working on the analysis, you can ignore this warning. When you’re finished, you can run wflow_publish to commit the R Markdown file and build the HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20190211) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Untracked files:
Untracked: alk_luad_synonymous.csv
Untracked: analysis/alk_luad_mutation_bias.Rmd
Untracked: analysis/mutation_bias_gist_brca.Rmd
Untracked: bias.brca.csv
Untracked: bias.gist.csv
Untracked: code/alldata_compiler.R
Untracked: code/contab_maker.R
Untracked: code/mut_excl_genes_datapoints.R
Untracked: code/mut_excl_genes_generator.R
Untracked: code/quadratic_solver.R
Untracked: code/simresults_generator.R
Untracked: data/ALKATI_ccle.csv
Untracked: data/All_Data_V2.csv
Untracked: data/CCLE_NP24.2009_Drug_data_2015.02.24.csv
Untracked: data/alkati_growthcurvedata.csv
Untracked: data/alkati_growthcurvedata_popdoublings.csv
Untracked: data/alkati_melanoma_vemurafenib_figure_data.csv
Untracked: data/alkati_simulations_compiled_1000_12319.csv
Untracked: data/all_data.csv
Untracked: data/gist_expression/
Untracked: data/tcga_brca_expression/
Untracked: data/tcga_luad_expression/
Untracked: data/tcga_skcm_expression/
Untracked: docs/figure/Filteranalysis.Rmd/
Untracked: mutation_bias_brca_n.pdf
Untracked: mutation_bias_brca_probabilities.pdf
Untracked: mutation_bias_brca_probabilities2.pdf
Untracked: mutation_bias_gist.pdf
Untracked: mutation_bias_gist_n.pdf
Untracked: mutation_bias_gist_probabilities.pdf
Untracked: mutation_bias_gist_probabilities2.pdf
Untracked: mutation_bias_luad.pdf
Untracked: mutation_bias_luad_deviations.pdf
Untracked: mutation_bias_luad_diagnostic.pdf
Untracked: mutation_probability.csv
Untracked: output/ alkati_subsamplesize_orval_fig1c.pdf
Untracked: output/alkati_ccle_tae684_plot.pdf
Untracked: output/alkati_filtercutoff_allfilters.csv
Untracked: output/alkati_luad_exonimbalance.pdf
Untracked: output/alkati_mtn_pval_fig2B.pdf
Untracked: output/alkati_skcm_exonimbalance.pdf
Untracked: output/alkati_subsamplesize_pval_fig.pdf
Untracked: output/alkati_subsamplesize_pval_fig1c.pdf
Untracked: output/all_data_luad.csv
Untracked: output/all_data_luad_egfr.csv
Untracked: output/all_data_skcm.csv
Untracked: output/baf3_alkati_figure_deltaadjusted_doublings.pdf
Untracked: output/baf3_barplot.pdf
Untracked: output/baf3_elisa_barplot.pdf
Untracked: output/egfr_luad_exonimbalance.pdf
Untracked: output/fig1c_3719_4.pdf
Untracked: output/fig2b2_filtercutoff_atinras_totalalk.pdf
Untracked: output/fig2b_filtercutoff_atibraf.pdf
Untracked: output/fig2b_filtercutoff_atinras.pdf
Untracked: output/luad_alk_exon_expression.csv
Untracked: output/luad_egfr_exon_expression.csv
Untracked: output/melanoma_vemurafenib_fig.pdf
Untracked: output/skcm_alk_exon_expression.csv
Untracked: output/suppfig1..pdf
Untracked: suppfig1..pdf
Unstaged changes:
Modified: analysis/ALKATI_Filter_Cutoff_Analysis.Rmd
Modified: analysis/ALK_ExonImbalance_SKCM_Analysis.Rmd
Modified: analysis/TCGA_luad_data_parser.Rmd
Modified: analysis/alkati_cell_line_tae684_response.Rmd
Modified: analysis/alkati_subsampling_simulations.Rmd
Modified: analysis/baf3_alkati_transformations.Rmd
Modified: analysis/index.Rmd
Modified: analysis/pairwise_comparisons_conditional_selection_simulated_cohorts.Rmd
Modified: analysis/practice.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the R Markdown and HTML files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view them.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | 6744c50 | haiderinam | 2019-03-06 | Build site. |
| html | aff917b | haiderinam | 2019-02-20 | Build site. |
| html | 0b5f5cb | haiderinam | 2019-02-19 | Build site. |
| html | 4b082e4 | haiderinam | 2019-02-19 | Build site. |
| html | 08e9438 | haiderinam | 2019-02-17 | Build site. |
| html | dfdb600 | haiderinam | 2019-02-17 | Build site. |
| Rmd | f8e2d5e | haiderinam | 2019-02-17 | Published Analysis on ALK expression levels #2 |
library(knitr)
library(tictoc)
library(workflowr)This is workflowr version 1.3.0
Run ?workflowr for help getting startedlibrary(VennDiagram)Loading required package: gridLoading required package: futile.loggerlibrary(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionlibrary(foreach)
library(doParallel)Loading required package: iteratorsLoading required package: parallellibrary(ggplot2)
library(reshape2)
library(RColorBrewer)
library(devtools)
library(ggsignif)
source("code/contab_maker.R")
source("code/alldata_compiler.R")
source("code/quadratic_solver.R")
source("code/mut_excl_genes_generator.R")
source("code/mut_excl_genes_datapoints.R")
source("code/simresults_generator.R")
######################Cleanup for GGPlot2#########################################
cleanup=theme_bw() +
theme(plot.title = element_text(hjust=.5),
panel.grid.major = element_blank(),
panel.grid.major.y = element_blank(),
panel.background = element_blank(),
axis.line = element_line(color = "black"))Making ALK Expression the plots:
alkati_merged_data=read.csv("output/all_data_skcm.csv")
alkati_merged_data$alkati=0
alkati_merged_data$alkati[alkati_merged_data$Ratio>=10&alkati_merged_data$mRNA_count>=500&alkati_merged_data$RSEM_normalized>=100]=1
alkati_merged_data$alkati=factor(alkati_merged_data$alkati,levels=c("1","0"))
ggplot(alkati_merged_data,aes(x=mean_RPKM_1.19, y=mean_RPKM_20.29,color=factor(alkati)))+
geom_abline(size=1)+
geom_point(size=4)+
####Had to add this line to not overplot the alkati datapoint- Haider 1/31/19
geom_point(data=alkati_merged_data[alkati_merged_data$alkati==1,],aes(x=mean_RPKM_1.19, y=mean_RPKM_20.29,color=factor(alkati)),size=4)+
scale_x_continuous(trans = "log10",name="Exon 1:19 RPKM",breaks=c(1e-2,1e0,1e2),labels = parse(text = c("10^-2","10^0","10^2")),limits = c(1e-3,1e3))+
scale_y_continuous(trans = "log10",name="Exon 20:29 RPKM",breaks=c(1e-2,1e0,1e2),labels = parse(text = c("10^-2","10^0","10^2")),limits = c(1e-3,1e3))+
scale_color_brewer(palette="Set1",name="ALKATI",labels=c("Yes", "No"))+
cleanup+
theme(plot.title = element_text(hjust=.5),
text = element_text(size=11,face = "bold"),
axis.title = element_text(face="bold",size="11"),
axis.text=element_text(face="bold",size="11",colour = "black"))+
theme(legend.key.size = unit(30,"pt"))Warning: Transformation introduced infinite values in continuous x-axisWarning: Removed 2 rows containing missing values (geom_point).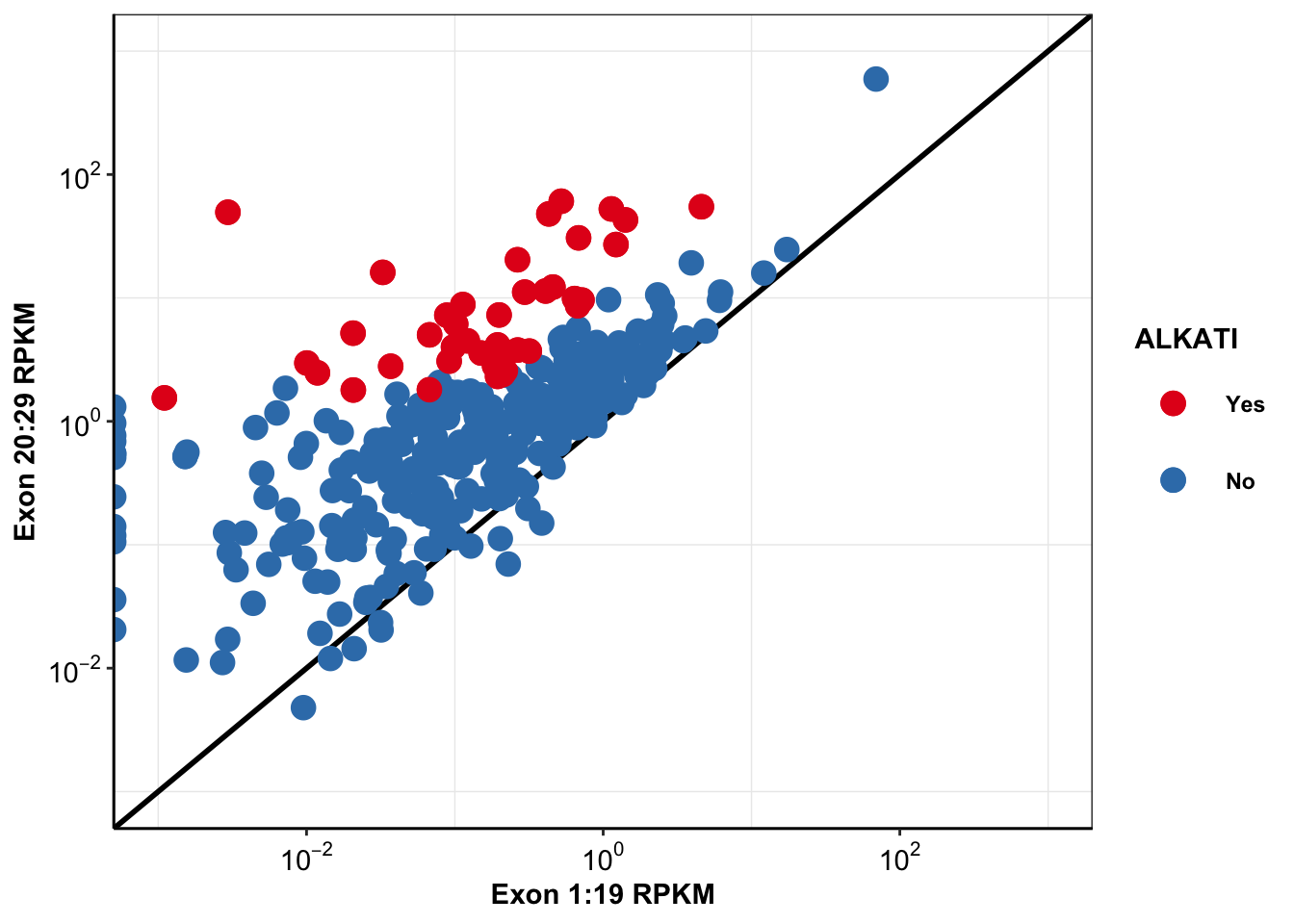
| Version | Author | Date |
|---|---|---|
| dfdb600 | haiderinam | 2019-02-17 |
# ggsave("output/alkati_skcm_exonimbalance.pdf",width =12 ,height =10 ,units = "in",useDingbats=F)Statistical test to see if ALK kinase domain expression was significantly higher than other domains
#Testing if both kinase and ALK expression are different
ks.test(alkati_merged_data$mean_RPKM_1.19,alkati_merged_data$mean_RPKM_20.29)Warning in ks.test(alkati_merged_data$mean_RPKM_1.19,
alkati_merged_data$mean_RPKM_20.29): p-value will be approximate in the
presence of ties
Two-sample Kolmogorov-Smirnov test
data: alkati_merged_data$mean_RPKM_1.19 and alkati_merged_data$mean_RPKM_20.29
D = 0.40456, p-value < 2.2e-16
alternative hypothesis: two-sided###We observed a significant difference between the distribution for the 20-29 exons and the 1-19 exons The reported p-value was 2-16.
###The p-value from a Chi-sq test was 2.2e-16 too
#for all ALK data, not ALKATI
ov_expr_obs=c(sum(as.numeric(alkati_merged_data$Ratio20.29>1),na.rm = T),
dim(alkati_merged_data)[1]-sum(as.numeric(alkati_merged_data$Ratio20.29>1),na.rm = T))
ov_expr_expected=c(round(dim(alkati_merged_data)[1]/2),
round(dim(alkati_merged_data)[1]/2))
overexpression=data.frame(rbind(ov_expr_obs,ov_expr_expected))
# overexpression=data.frame(rbind(c(338,13),c(176,176)))
colnames(overexpression)=c("Yes","No")
chisq.test(overexpression)
Pearson's Chi-squared test with Yates' continuity correction
data: overexpression
X-squared = 189.29, df = 1, p-value < 2.2e-16
sessionInfo()R version 3.5.2 (2018-12-20)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Mojave 10.14.5
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] parallel grid stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] ggsignif_0.5.0 usethis_1.5.0 devtools_2.0.2
[4] RColorBrewer_1.1-2 reshape2_1.4.3 ggplot2_3.1.1
[7] doParallel_1.0.14 iterators_1.0.10 foreach_1.4.4
[10] dplyr_0.8.1 VennDiagram_1.6.20 futile.logger_1.4.3
[13] workflowr_1.3.0 tictoc_1.0 knitr_1.23
loaded via a namespace (and not attached):
[1] tidyselect_0.2.5 xfun_0.7 remotes_2.0.4
[4] purrr_0.3.2 colorspace_1.4-1 htmltools_0.3.6
[7] yaml_2.2.0 rlang_0.3.4 pkgbuild_1.0.3
[10] pillar_1.4.0 glue_1.3.1 withr_2.1.2
[13] lambda.r_1.2.3 sessioninfo_1.1.1 plyr_1.8.4
[16] stringr_1.4.0 munsell_0.5.0 gtable_0.3.0
[19] codetools_0.2-16 evaluate_0.13 memoise_1.1.0
[22] callr_3.2.0 ps_1.3.0 Rcpp_1.0.1
[25] scales_1.0.0 backports_1.1.4 formatR_1.6
[28] desc_1.2.0 pkgload_1.0.2 fs_1.3.1
[31] digest_0.6.19 stringi_1.4.3 processx_3.3.1
[34] rprojroot_1.3-2 cli_1.1.0 tools_3.5.2
[37] magrittr_1.5 lazyeval_0.2.2 tibble_2.1.1
[40] futile.options_1.0.1 crayon_1.3.4 whisker_0.3-2
[43] pkgconfig_2.0.2 prettyunits_1.0.2 assertthat_0.2.1
[46] rmarkdown_1.12 R6_2.4.0 git2r_0.25.2
[49] compiler_3.5.2